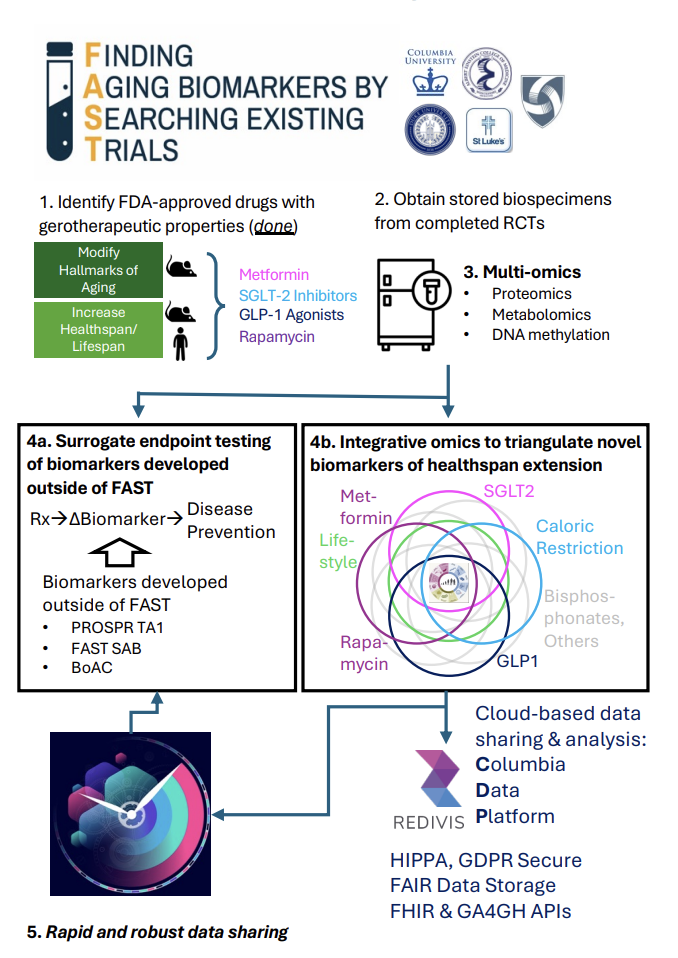
Finding Aging Biomarkers by Searching Existing Trials (FAST)
Aging is the leading cause of most chronic diseases driving population burden of disability and premature mortality. The decline our bodies experience as we grow older is accepted as normal. It shouldn’t be. We can see around us the examples of successful aging; people live through their 80s and 90s cognitively intact and in good health. Breakthroughs in geroscience establish the biology of aging can be modified; interventions in laboratory animals that slow biological processes of aging increase healthspan. These preclinical discoveries suggest that, for humans, maintaining health and function into the 9th decade of life and beyond is possible for all of us. FAST is motivated by the long-term goal of enabling more people to realize this potential for healthspan extension. To achieve this goal, we aim to move biomedical research away from geriatrics-clinic whack-a-mole, in which we develop treatments for symptomatic diseases one at a time, and toward therapies that maintain health and prevent disease. Preclinical studies are bringing these therapies within reach. But translating them in human trials faces the challenge that, unlike in short-lived laboratory models, determining impacts on human healthspan will require decades of follow-up. To overcome this challenge, biomarkers are needed that are responsive to intervention over timescales of months-to-years and which can provide surrogates for the long-term impacts of novel therapies on healthspan. The immediate objective of FAST-PROSPR is to establish sensitive and specific biomarkers that can be tested to determine whether an intervention is slowing the biology of aging and extending healthspan.
Trials included in FAST:
FAST has enrolled 6 trials: The Diabetes Prevention Program (DPP) tested metformin and lifestyle to prevent incident diabetes. PRESERVED-HF and DEFINE-HF tested the SGLT-2 Inhibitor dapagliflozin to protect function and healthspan in people with heart failure. SELECT tested the GLP1 agonist semaglutide to prevent cardiovascular events in people with cardiovascular disease and overweight. VIBRANT tested rapamycin to promote ovarian reserve in healthy young women. CALERIE tested 25% caloric restriction to reduce resting metabolic rate and core temperature.
Collectively, these trials provide access to thousands of samples from successful trials that establish intervention effects on healthspan-relevant endpoints. We are continuing to explore opportunities with Novo Nordisk and Astra Zeneca to access additional GLP1 trial data.
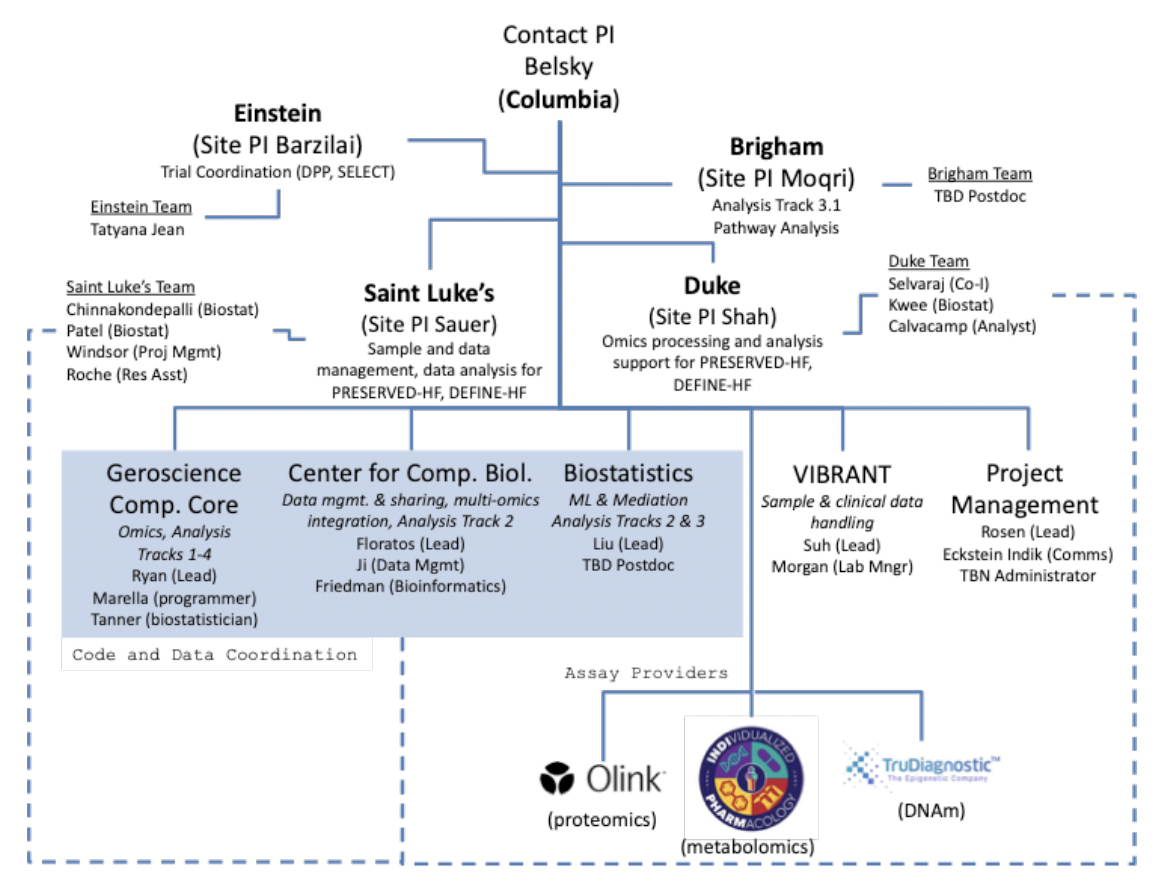
Meet the Team.
Our project brings together teams from five institutions with specific expertise in aging biology and biomarkers of aging, clinical trials, biospecimen handling, proteomics, metabolomics, epigenetics, biostatistics, bioinformatics, and computational biology, and data management and sharing.
Project Leadership
Dan Belsky, PhD is Associate Professor of Epidemiology at the Columbia University Mailman School of Public in the Robert N Butler Columbia Aging Center, where he directs the Center’s Geroscience Computational Core. His group develops methods to quantify the pace and progress of biological aging in young, midlife, and older adult humans and applies these methods in epidemiological studies and clinical trials to identify opportunities for intervention to increase healthy lifespan. Dan has been recognized as a leader in his field with the Academy of Behavioral Medicine’s Neal Miller New Investigator Award and the American Federation for Aging Research’s Vincent Cristofalo Rising Star Award. His work is supported by the US National Institute on Aging, the Advanced Research Projects Agency for Health (ARPA-H), the Canadian Institute for Advanced Research, and the American Federation for Aging Research (AFAR), among other sources. He is co-director of the AFAR FAST Initiative and a member of the Scientific Advisory Boards of AFAR, the Biomarkers of Aging Consortium, and X-Prize Healthspan. He is an inventor of the Pace of Aging method and the DunedinPACE epigenetic clock. Since 2020, he has been named an ISI highly-cited researcher.
Nir Barzilai, MD, is a leading figure in the field of geroscience, whose pioneering work has shown that aging itself has distinct biological drivers that can be targeted to prevent age-related diseases. At Albert Einstein College of Medicine, he serves as Professor of Medicine and Genetics and directs the Institute for Aging Research and BIO VITAL, the Center for Validation of Interventions Targeting Aging and Longevity. Dr. Barzilai has published more than 360 papers and made seminal contributions to understanding the mechanisms of exceptional longevity, extending healthspan in model organisms, and identifying genetic pathways linked to healthy aging in centenarian families. He is the recipient of numerous honors, including the IPSEN Longevity Award. He co-founded and serves as President of the Academy of Geroscience and sits on the Board of Directors of the American Federation for Aging Research, where he co-leads the biomarker program (FAST, ARPA-H), the Targeting Aging with Metformin (TAME) trial, and the Super Agers initiative. He also serves on the Executive Committee of the Longevity Biotech Association, the Council of the Healthy Longevity Medicine Society, and is a Fellow of both the Association of American Physicians (AAP) and the New York Academy of Medicine. Learn more about Dr. Barzilai’s work at https://einsteinmed.edu/centers/aging/.
-
Tatyana Jean-Davis, MPH is an experienced clinical research professional with over 15 years of experience leading and managing complex clinical trials across academic medical centers, biotech organizations, and global CRO environments. Throughout her career, she has consistently managed multifaceted clinical programs involving high-volume data collection, biospecimen oversight, and cross-functional team leadership. Much of her work has centered on therapeutic areas deeply relevant to geroscience and biomarker discovery, including oncology, infectious disease, gastroenterology, and nuclear medicine. These experiences have given her a unique appreciation for the operational, scientific, and logistical complexities involved in large-scale biomarker evaluation efforts like FAST. Tatyana holds a Master of Public Health from Thomas Jefferson University and a Bachelor of Arts in Psychology from the University at Albany, SUNY.
Mahdi Moqri, PhD, MBA, is a faculty member at Harvard Medical School and an independent investigator at the Division of Genetics as well as the Endocrinology, Diabetes, and Metabolism. His research integrates omics, bioinformatics, and computational methods to uncover the molecular mechanisms of aging and to develop new approaches for early detection and prevention of age-related diseases. Dr. Moqri co-directs the Biomarkers of Aging Consortium (agingconsortium.org), where he leads efforts to develop blood-based biomarkers of aging using large-scale omics datasets and longitudinal biobank resources. He completed his postdoctoral fellowship at Stanford University in the Snyder Lab, and his formal training is in computer science and informatics. Learn more about Dr. Moqri’s work at https://pepper.bwh.harvard.edu/.
Robert N. Butler Columbia Aging Center
FAST is based in the Robert N Butler Columbia Aging Center, where PI Belsky directs the Center's Geroscience Computational Core.
Zohn Rosen, PhD, Project Management Lead, is an Assistant Professor of Health Policy and Management at the Columbia University Mailman School of Public Health. Dr. Rosen is an experimental psychologist, with an extensive history of conducting large and complex studies in the fields of health behaviors and social epidemiology, as well as projects that include biological markers of health and aging. Zohn will lead project management lead for the ARPA-H FAST initiative.
-
Rosine Moussa, MA is the Center Administrator at the Robert N. Butler Columbia Aging Center, a role she has held since 2016. With more than 20 years of progressive administrative experience at Columbia University, she brings deep expertise in research administration, operations, and compliance. Her work focuses on supporting complex, multidisciplinary research initiatives by overseeing administrative strategy, financial management, human resources, and regulatory compliance to ensure the effective execution of sponsored projects.
-
Claire Eckstein Indik serves as a Project Coordinator at the Robert N. Butler Columbia Aging Center at Columbia University. In this role, she supports project management, coordination, and communications for the ARPA-H FAST initiative.
Robert N. Butler Columbia Aging Center: Geroscience Computational Core
Calen Ryan, PhD, Geroscience Computational Core Data Science Lead, is an evolutionary biologist and human geneticist specializing in the role of epigenetic processes in development, reproduction, aging, and health. He works with high-throughput data from large, longitudinal population and clinical datasets to answer fundamental questions about why our bodies function the way they do, and how that can help us be healthier, and age better. Dr. Ryan’s overarching research goal is to use an evolutionary framework to inform basic biology, medicine, and public health by characterizing the molecular processes underlying evolutionarily-motivated questions. He is most interested in reproduction, inheritance, development, and senescence (aging), which are core concepts in evolutionary thinking. Learn more about Dr. Ryan’s work at https://cpryan.github.io/
-
Will Marella, MPhil, is a Senior Data Analyst in the Robert N. Butler Columbia Aging Center. He received a BA in Biology from Brown University in 2023 and an MPhil in Population Health Sciences from the University of Cambridge in 2024. For his MPhil dissertation, he mathematically derived, developed, and analyzed a mean field variational inference algorithm and partially collapsed Gibbs sampler to optimize probabilistic modeling used to identify multimorbidities from health data. In the Belsky lab, Will is a member of the Geroscience Computational Core, where he develops key GCC packages and applies machine learning and statistical modeling to complex data types to assist a variety of aging-related research projects. Learn more about Will’s work at https://github.com/will-marella
Department of Biostatistics
Zhonghua Liu, ScD, is an Assistant Professor of Biostatistics at the Mailman School of Public Health and a member of the Data Science Institute at Columbia University. He is a biostatistician specializing in statistical genetics, genomics, and bioinformatics, with core expertise in causal inference and machine (deep) learning methods for large-scale genetic and multi-omics data. His research focuses on developing robust methods for Mendelian randomization, causal mediation analysis, and semiparametric and machine learning–based inference to elucidate molecular mechanisms and causal pathways underlying complex traits and diseases, with particular emphasis on aging and GeroScience. He serves as an Associate Editor for the Journal of Statistical Planning and Inference, Statistics in Medicine and is on the editorial board of BMC Bioinformatics, and regularly reviews for leading journals in statistics, genetics, and biomedical research. Learn more about Dr. Liu’s work at https://sites.google.com/view/drliu/home.
Departments of Obstetrics & Gynecology and Genetics & Development
Yousin Suh, PhD, is the Charles and Marie Robertson Professor of Reproductive Sciences in Obstetrics and Gynecology, Professor of Genetics and Development, and Director of Reproductive Aging in Obstetrics and Gynecology in the Vagelos College of Physicians and Surgeons at Columbia University. She investigates the (epi)genetic component that underlies the interface of intrinsic aging and disease based on the identification of (epi)genome sequence variants associated with age-related disease risk or its opposite, i.e., an unusual resistance to such disease. To investigate the functional impact of (epi)genetic variants, she applies specific functional tests, including in silico modeling, cell culture assays and mouse models. Her contributions in the field have been recognized by the Glenn Award for Research in Biological Mechanisms of Aging and has led to become the elected member of the Academy for Health and Lifespan Research, the first global non-profit group focused on accelerating breakthroughs in the expansion of the human health span, nominated and voted on by the world’s leading geroscientists. She has organized numerous international symposiums on functional genomics of aging, is on the Editorial Boards of numerous Journals including PLoS Genetics and Aging Cell as an Associate Editor, and participates in advisory committee members for several research institutions and companies. Learn more about Dr. Suh’s work at http://yousinsuhlab.org/.
Columbia University Department of Systems Biology: Center for Computational Biology and Bioinformatics
Aris Floratos, PhD, Center for Computational Biology and Bioinformatics Lead, is an Associate Professor in the Departments of Systems Biology and Biomedical Informatics, Executive Research Director of the Center for Computational Biology and Bioinformatics, and Scientific Director of the Bioinformatics Core at the Herbert Irving Comprehensive Cancer Center. A computer scientist by training, his research focuses on bioinformatics, algorithms, and computational theory applied to problems in human health. Dr. Floratos’s group develops innovative, systems biology–driven methods to improve the detection of low-risk genetic factors contributing to serious drug-induced adverse events. Working with an international network of collaborators, his lab has led large-scale analyses of genome-wide genotyping and exome sequencing data for drug-induced conditions including severe cutaneous reactions, liver injury, cardiac arrhythmias, and osteonecrosis of the jaw. His group also has a strong track record in building software tools and data platforms for the analysis, management, and dissemination of genomic data, including portals that support data submission and sharing for large public and private research consortia. Learn more about Dr. Floratos’s work at https://systemsbiology.columbia.edu/faculty/aris-floratos.
-
Richard Friedman PhD, is Associate Research Scientist in the Herbert Irving Comprehensive Cancer Center, where he serves as a consultant in the Bioinformatics Core, and Lecturer in the Department of Biomedical Informatics, Columbia University Irving Medical Center. Dr. Friedman analyzes transcriptomic, epigenetic, and metabolomic data to characterize the molecular basis of cancer, cardiovascular disease, and ageing. Dr. Friedman was trained as physical chemist. Prior to his work in bioinformatics, he developed theories and performed computations on the stability and reactions of biological molecules. Learn more about Dr. Friedman’s work at https://www.columbia.edu/~raf4/index.html.
-
Zhou Ji, PhD, is a Lead Programmer Analyst in the Department of Systems Biology. He leads complex software projects and conducts cross-disciplinary research in biological sciences, data science, and software engineering. Learn more about Dr. Ji’s publications at Google Scholar.
Gary W. Miller, PhD is the Adrienne Block Professor of Environmental Health Sciences (in Molecular Pharmacology and Therapeutics), Vice Dean for Research Strategy and Innovation at the Mailman School of Public Health, Co-Director, Precision Medicine Core, Irving Institute and founding director of the Center for Innovative Exposomics. Dr Miller is a leader in the exposome field, which strives to provide a systematic and comprehensive analysis of the non-genetic contributors to health and disease. Currently, he leads the Network for Innovative Exposomics (NEXUS) and Individual metabolome and exposome assessment for Pharmaceutical optimization (IndiPHARM). Dr Miller was the founding director of the HERCULES Exposome Research Center at Emory University, the first exposome-based research center in the U.S. He authored the first book on the topic, The Exposome: A Primer published by Elsevier. His research focuses on environmental drivers of neurodegeneration. His laboratory uses a variety of methods including transgenic mouse production, immunohistochemistry, neurotransmitter transport assays, high-resolution metabolomics, electrochemistry, and behavioral assays. His work is conducted in several experimental models from cultured neurons and C. elegans to mice and human studies. Learn more about the Center for Innovative Exposomics.
-
Sophie Thuault-Restituito, PhD serves as the Executive Director of Strategic Initiatives, directly reporting to the funding Director, Gary Miller. In this capacity, Sophie works closely with Gary to establish and implement the Center's strategic vision. Before undertaking this role, Sophie held the position of Chief of Staff and Executive Director for Special Projects for the the Executive Vice President for Research, Jeannette Wing. Preceding her tenure there, she assumed the role of Chief of Staff of the Irving Institute for Cancer Dynamics, working under Simon Tavare’s leadership to facilitate the institute's inception. Prior to IICD, Sophie spent two years as Executive Director at the Center for Topology of Cancer Evolution and Heterogeneity at CUIMC. Before joining Columbia University, Sophie was at NYU Langone Health for 13 years, first as a researcher in neuroscience and biochemistry, and later as operations manager/director for research facilities. Sophie received her doctorate in pharmacy and her PhD in neuroscience from the University of Montpellier, France, and completed a postdoc at the University of College London.
-
Randolph Reyes Singh, PhD is an Assistant Professor in the Department of Environmental Health Sciences at Columbia Mailman School of Public Health. His primary focus lies at the intersection of human health, environmental health, and analytical chemistry. Randolph uses high resolution mass spectrometry-based nontarget analysis (NTA) and orthogonal techniques to explore the chemical exposome in different human disease cohorts in order to help us understand the role of chemical exposures in disease development and progression. He is also interested in using NTA to highlight differences in chemical exposures arising from social disparities and inequities. Learn more about mass spectomerty experiments performed at the Biomarkers Core Laboratory (BCL).
Duke University
Svati H. Shah, MD, MHS, is a physician scientist and Associate Dean of Genomics and Director of Precision Genomics Collaboratory in the Duke School of Medicine; Vice-Chief of Translational Research and Director of the Adult Cardiovascular Genetics Clinic in the Division of Cardiology, Department of Medicine; Co-Director of Translational Research in the Duke Molecular Physiology Institute (DMPI); and a faculty member in the Duke Clinical Research Institute (DCRI). Her research focus is on metabolic and genetic pathways of cardiometabolic diseases, integrating diverse genomic, metabolomic and proteomic techniques for identification of novel mechanisms of disease and biomarkers. Her multi-disciplinary molecular epidemiology lab within the DMPI has quantitative and molecular components and leverages large biorepositories on to perform discovery studies using omics technologies, with subsequent functional validation for mechanistic insight.
-
Senthil Selvaraj, MD, MS, MA, is an Assistant Professor of Medicine in the Duke Molecular Physiology Institute and the Division of Cardiology at Duke University Medical Center. Dr. Selvaraj’s translational research explores the therapeutic relevance of cardiovascular metabolism to patients with heart failure. Through early phase work, his research employs deep phenotyping to decipher metabolic mechanisms that may be leveraged for cardiovascular benefit. These studies characterize dynamic changes in exercise physiology, biomarker profiles integrating multi-omic platforms, echocardiography, arterial stiffness, metabolic molecular imaging techniques, and several other modalities. More recently, this line of inquiry has explored the potential benefits of endogenous and exogenous ketogenic therapies among patients with heart failure. Dr. Selvaraj’s work is currently or recently funded by the National Institutes of Health, American Heart Association, Doris Duke Charitable Foundation, Mandel Foundation, Heart Center Leadership Council, Institute for Translational Medicine and Therapeutics, and American Society for Nuclear Cardiology. Learn more about Dr. Selvaraj’s work at https://dmpi.duke.edu/senthil-selvaraj-md-ms-ma.
Saint Luke’s Health System
Andrew J. Sauer, MD is a cardiologist and physician-scientist at Saint Luke’s Mid America Heart Institute (MAHI) in Kansas City and Associate Professor at the University of Missouri–Kansas City (UMKC). He serves as Medical Director of the Haverty Family Cardiometabolic Center of Excellence. Dr. Sauer’s clinical and research focus centers on heart failure, cardiometabolic disease, and device-based and digital strategies for hemodynamic optimization. He has served as a principal investigator and national leader across multiple multicenter trials evaluating pharmacologic, device, and remote monitoring therapies in heart failure, including pulmonary artery pressure monitoring, valvular interventions, and cardiometabolic agents. His work integrates guideline-directed medical therapy with real-time physiologic monitoring and precision phenotyping to improve outcomes in patients with heart failure across the ejection fraction spectrum. He has authored and co-authored numerous peer-reviewed publications and frequently speaks nationally on heart failure therapeutics, congestion management, cardiometabolic integration, and health system implementation science.